What's the LiverAtlas?
|
LiverAtlas is a integrated database of biomedical knowledge related to the liver and hepatic disease. It covers detail information on liver-related genome, liver transcriptome, liver proteome, liver metabolome, liver-related pathway and liver disease. LiverAtlas provides a wealth of manually curated information, relevant literature citations and cross-references to other databases. Especially, the database contains proteins and genes that specificly expressed in the liver.
How to get started with LiverAtlas?
|
Users can get started by browsing or searching the data in LiverAtlas.
(1) Data browse
Users can browse gene, protein, transcriptomic dataset, pathway and disease information though the Browse menu on the homepage.
(2) Data search
Users can search gene, protein, transcriptomic dataset, pathway and disease information though the searching bar on the left of the homepage.
For gene searches, users can use LiverAtlas Gene ID, Entrez Gene ID, official gene symbols, gene names, or synonyms. A similar search protocol is used for protein, transcriptomic dataset, pathway, or disease searches. Users can easily search for one term and be directed to all the related terms in the database.
What could I do with the data in LiverAtlas?
Users can find biomarkers based on the data in LiverAtlas. There are several examples of data analysis.
What's the scoring rule of the data reliability in LiverAtlas?
To facilitate users' selection of data reliability, liver-expressed genes and proteins, PPIs, PTMs, and molecular and genetic events of hepatic disease were assigned 5-grade reliability scores (RS), which were represented by five orange stars . These RSs were calculated using a semi-quantitative assessment, which considers both the reliability and the number of data sources at each reliability level. For each data item, a RS was obtained by applying the following algorithm:
. These RSs were calculated using a semi-quantitative assessment, which considers both the reliability and the number of data sources at each reliability level. For each data item, a RS was obtained by applying the following algorithm:
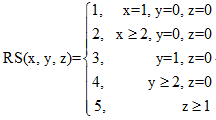
where x, y, and z represent respectively the number of data sources for each item of information in LiverAtlas. These data were originally derived from predictive studies, high-throughput experiments, and highly focused biochemical studies. Especially for molecular and genetic events in hepatic diseases, x, y, and z values might alternatively represent the number of observations acquired by automatic text mining by the computer, automatic text mining combined with manual analysis, and expert reading and confirmation. Corresponding RSs were calculated separately for each datum. Reliability for each item of information was classified according to RS value as low (RS<=2), medium (2<=RS<=4) and high (RS=5). This way, the user can choose to view high-confidence data alone or alternatively to reduce the search stringency and view additional data that have less confidence.
Where could I find more detailed help information of LiverAtlas?
The sitemap illustrates the detailed information of each kind of page in LiverAtlas and points out the links between them.
What's HuLDO?
Human Liver Disease Ontology (HuLDO) is a classification system of liver diseases. It contains definitions and descriptions of each kind of liver disease and points out their relations. HuLDO has been used as a catalogue to group the disease information in LiverAtlas, or as a dictionary to annotate the genes, proteins and pathways. You can browse disease information along its tree structure.
HuLDO can be downloaded freely from here and be used in further ways.
How could I cite LiverAtlas in my paper?
If you use LiverAtlas in any published work, please cite the following web address: http://liveratlas.hupo.org.cn/
How to contact us?
Postal Address: 33, Life Science Park Road, Changping District, Beijing, 102206, China
Telephone : 8610-80705225
Fax : 8610-80705155
E-mail: liveratlas@gmail.com